Parkin Agonist Development
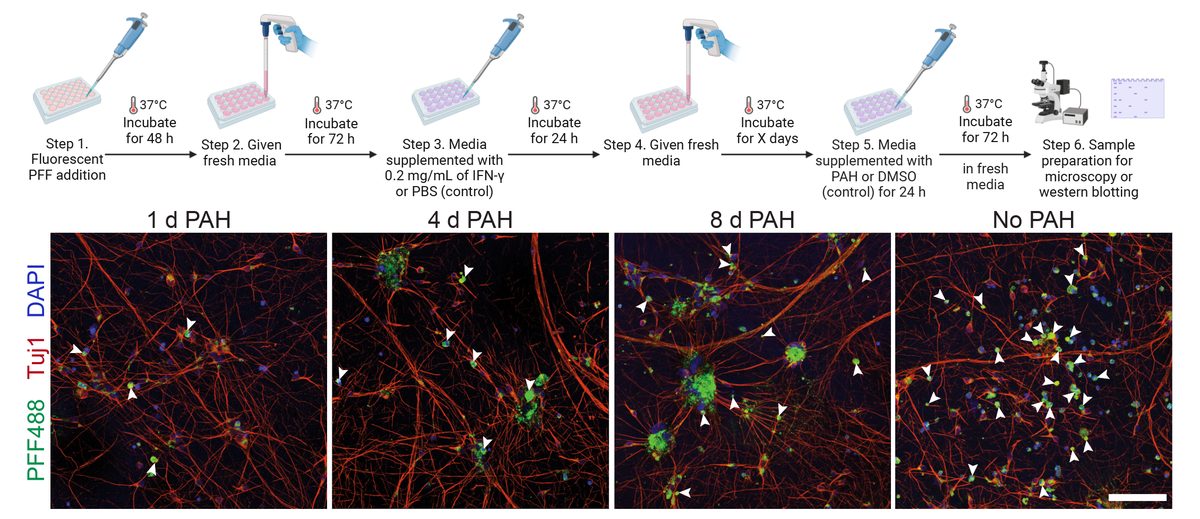
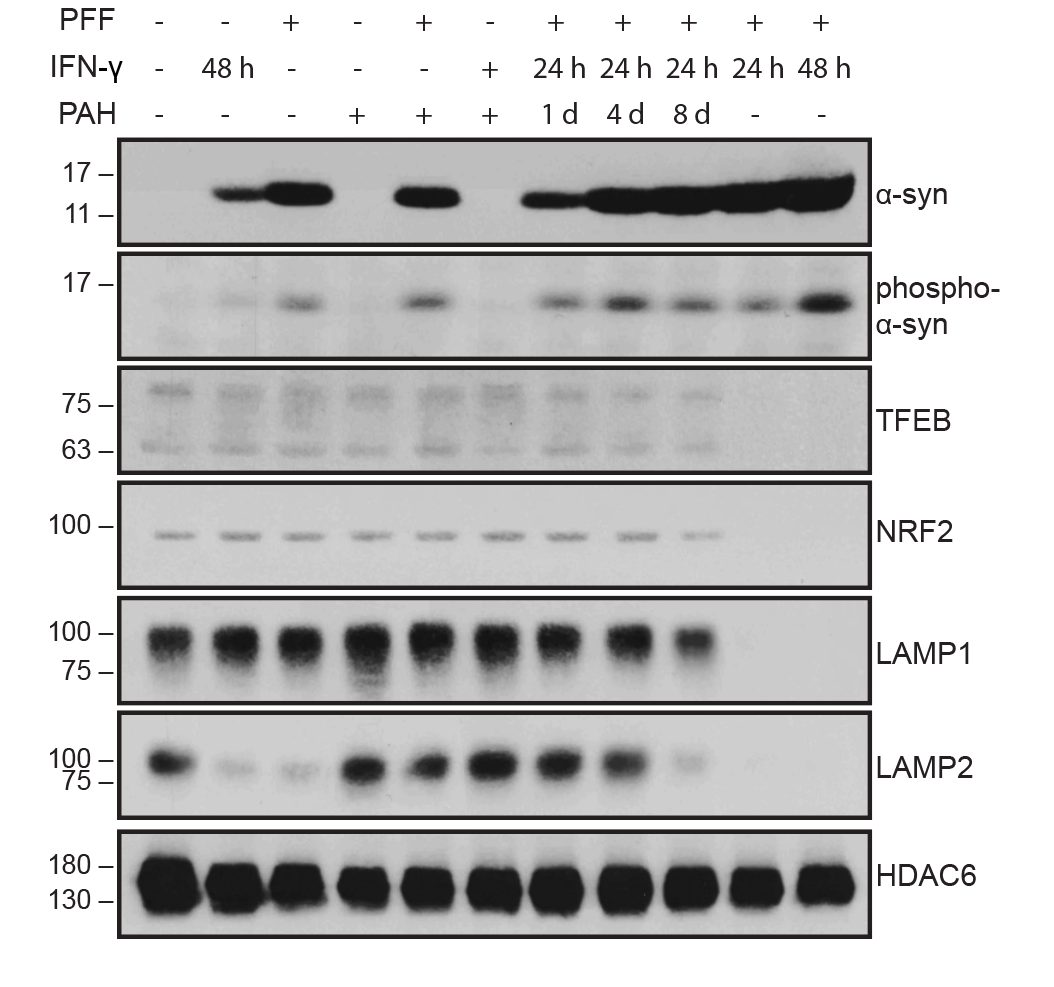
I collaborated with chemical biologists to identify small-molecule activators of Parkin. In dopaminergic neurons, Parkin agonists enhanced mitophagy, improved survival, and protected against α-synuclein–induced degeneration. I also demonstrated that Parkin loss shifts cell fate toward glial lineages, linking target perturbation to lineage-state biomarkers for translational stratification.
I generated oncology-facing rationale integrating observations that PARK2 downregulation is common across cancers with cell-based efficacy readouts. Manuscript under review, Nature.