iPSC Maintenance, Expansion & Differentiation
My iPSC culturing, maintenance, and differentiation training was done in association with the EDDU at McGill, generating some of the most homogenous cultures in the world. iPSCs from 9 different donors were used for modeling Parkinson's disease pathology. I have generated and banked over 20 different iPSC cell lines.
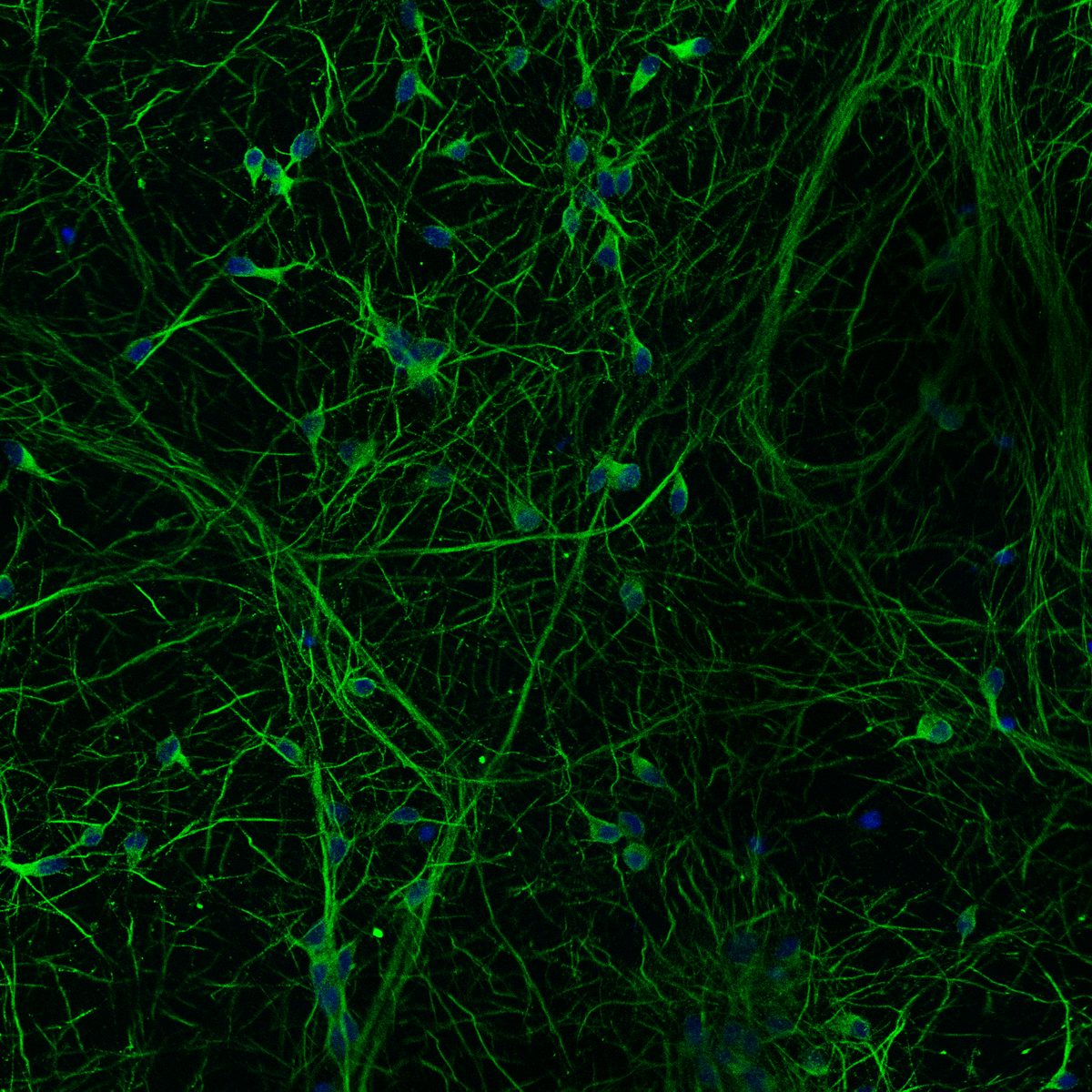
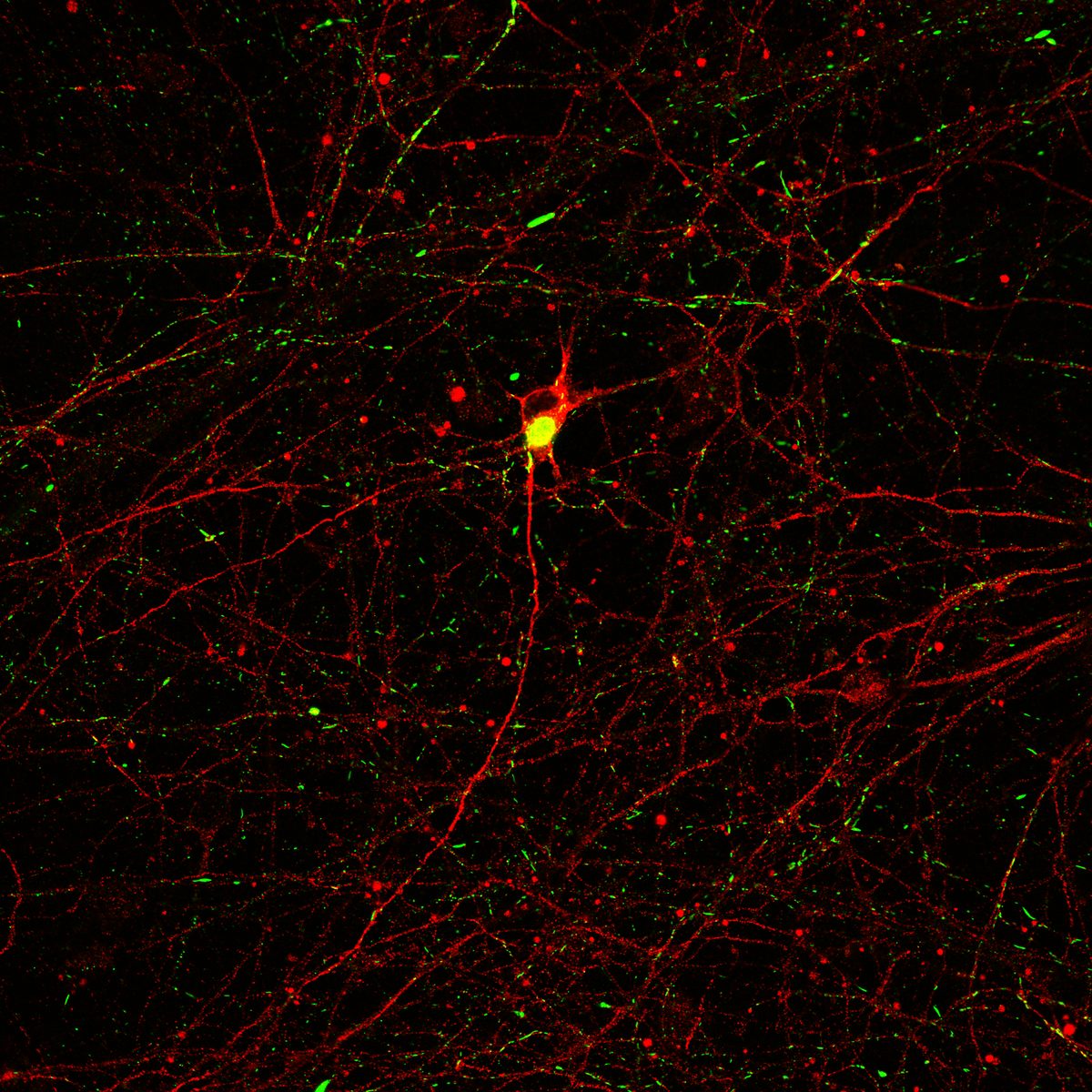
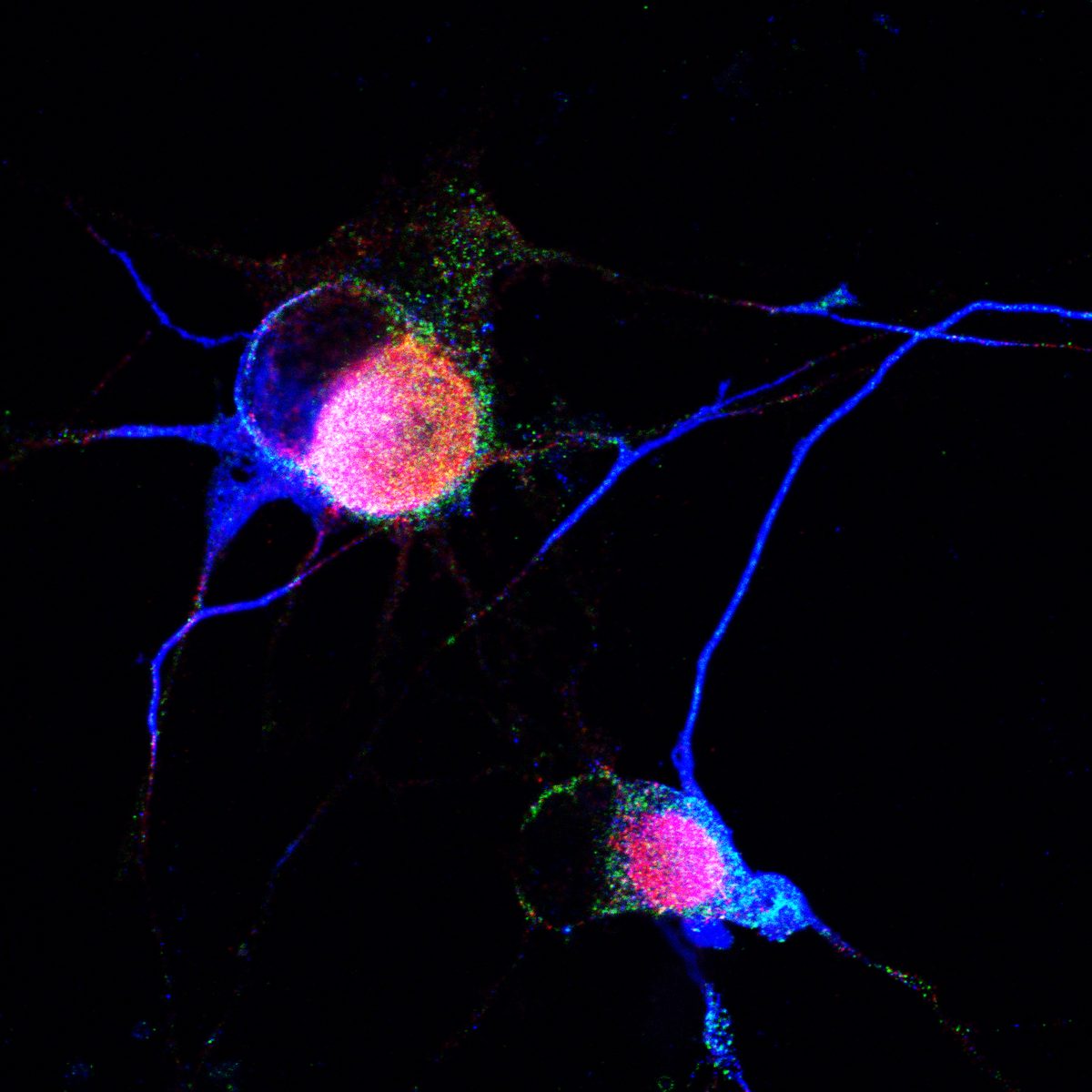
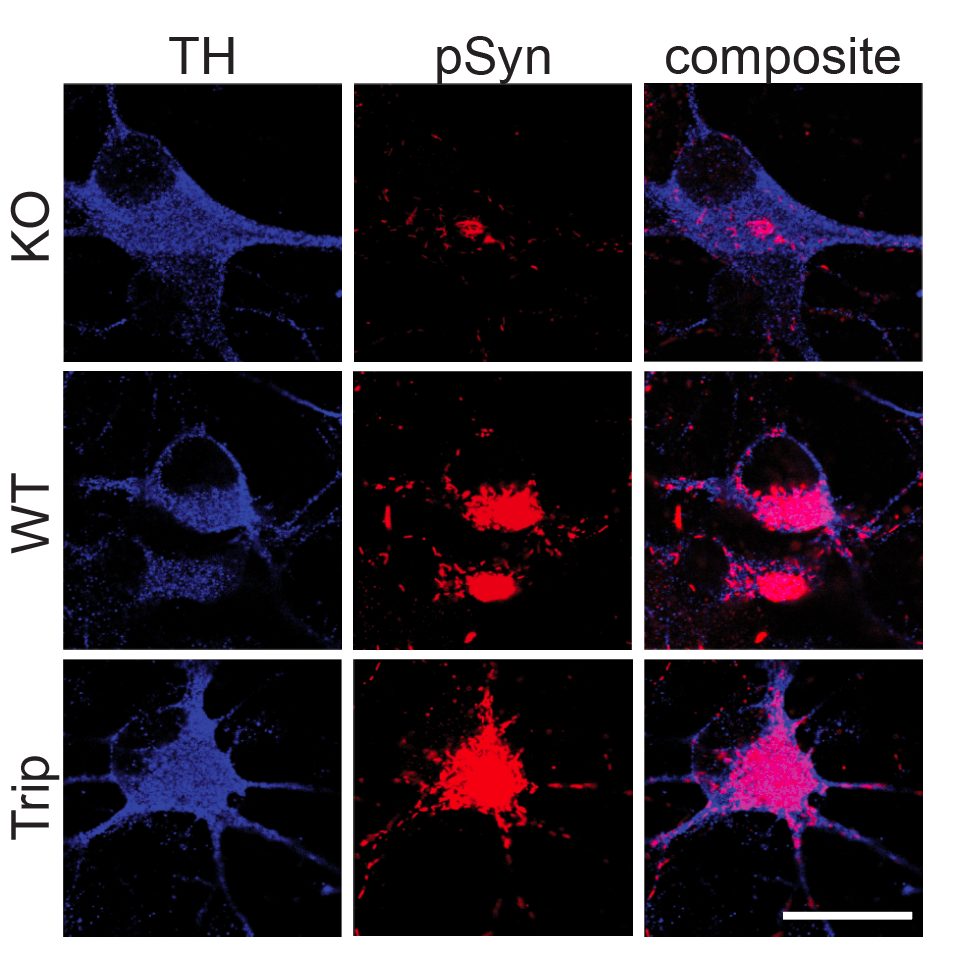
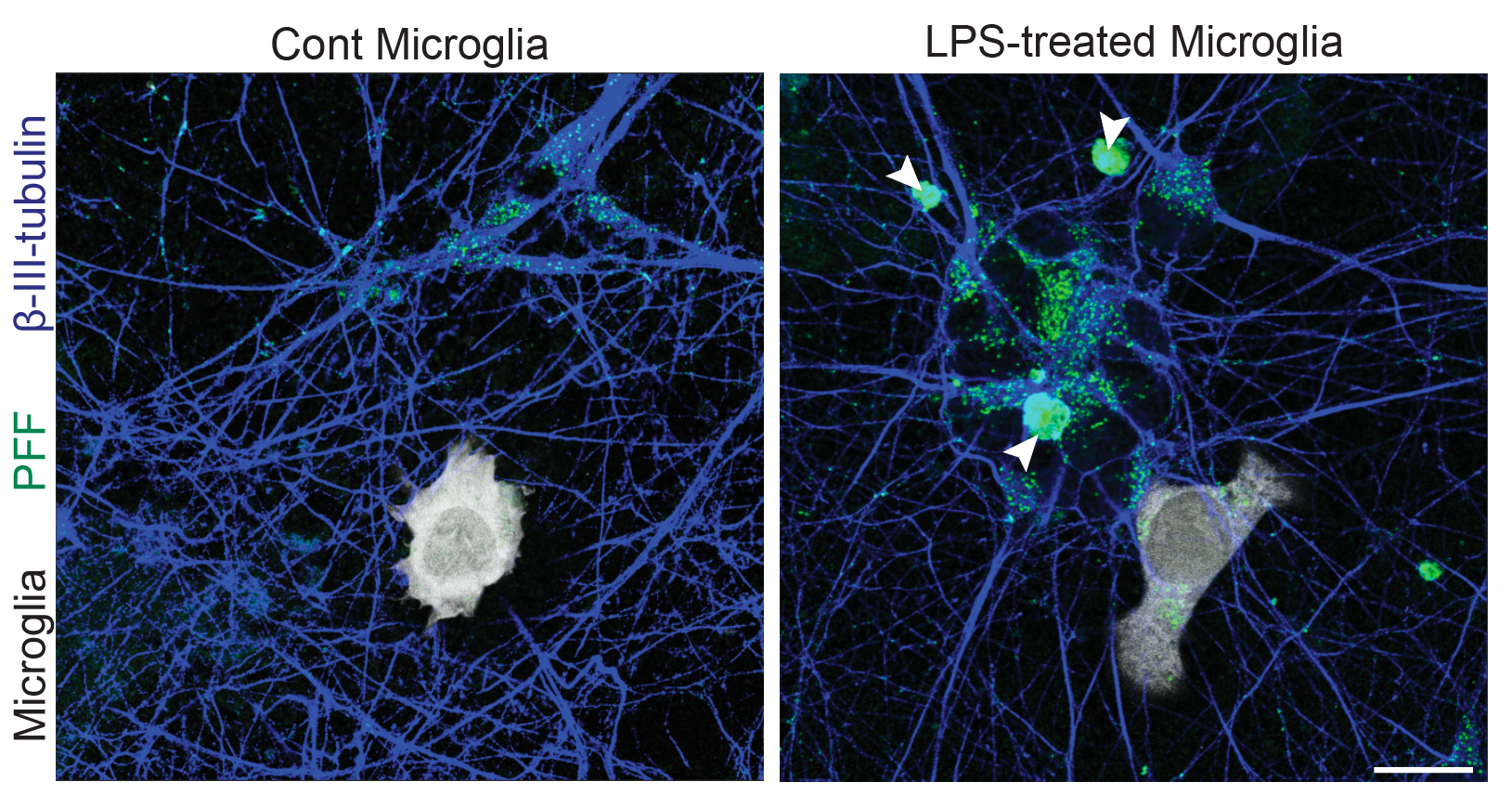
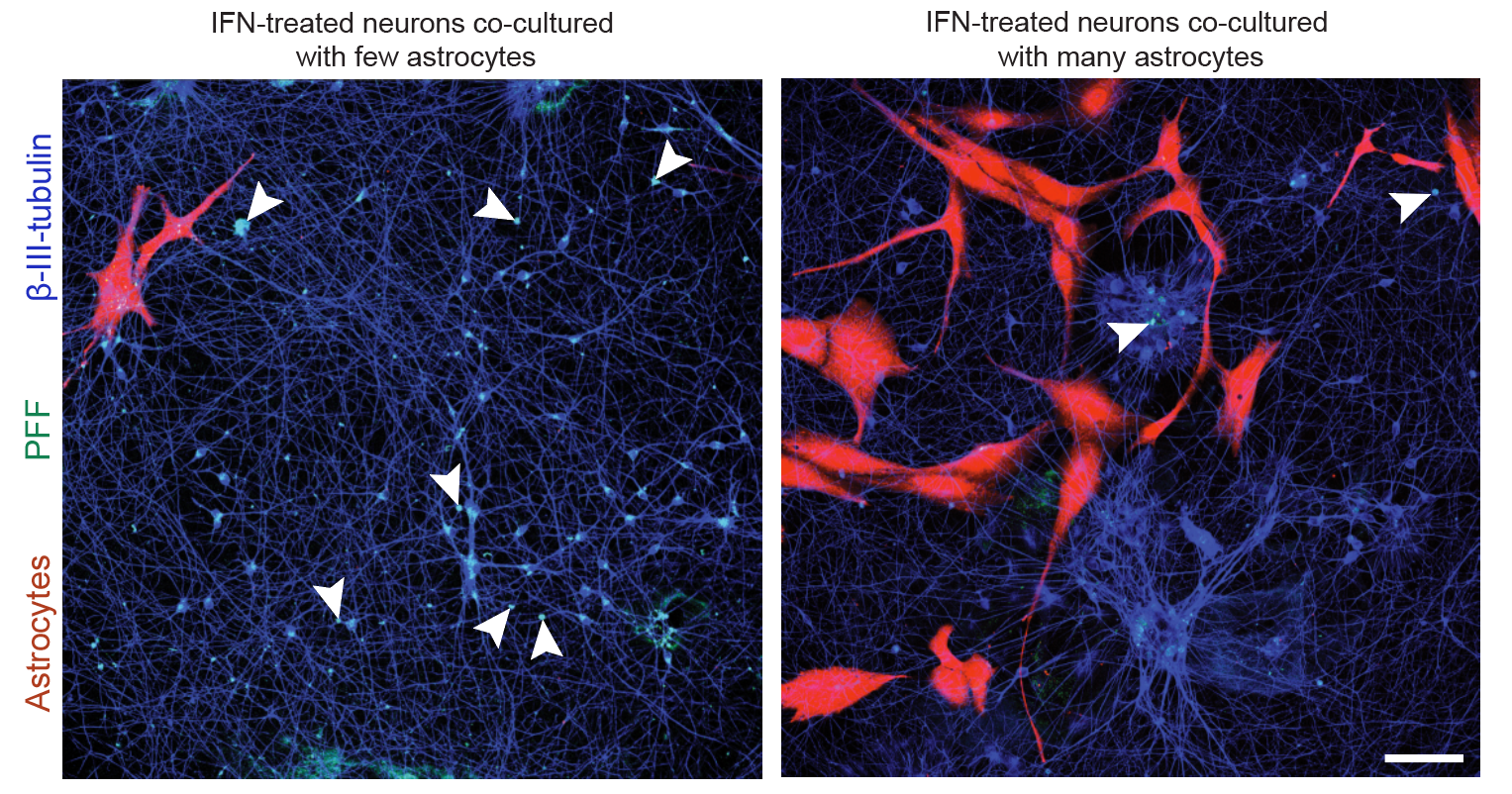
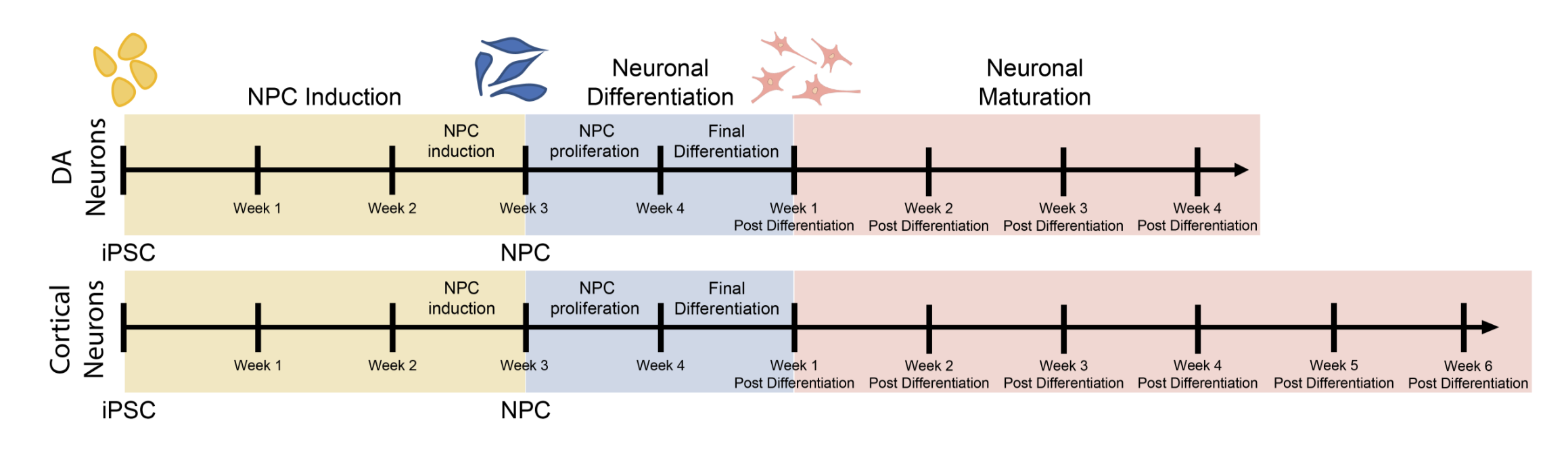
Differentiation spans ectodermal lineages (dopaminergic, cortical/excitatory neurons, astrocytes, microglia-like cells) and endodermal lineages (hepatic, lung/epithelial, gut, beta islet), in both adherent and suspension culture formats. Neuronal maturation requires 4–6 weeks to achieve full dopaminergic identity (DAT, GIRK2) or cortical maturity (VGLUT1, GABA).
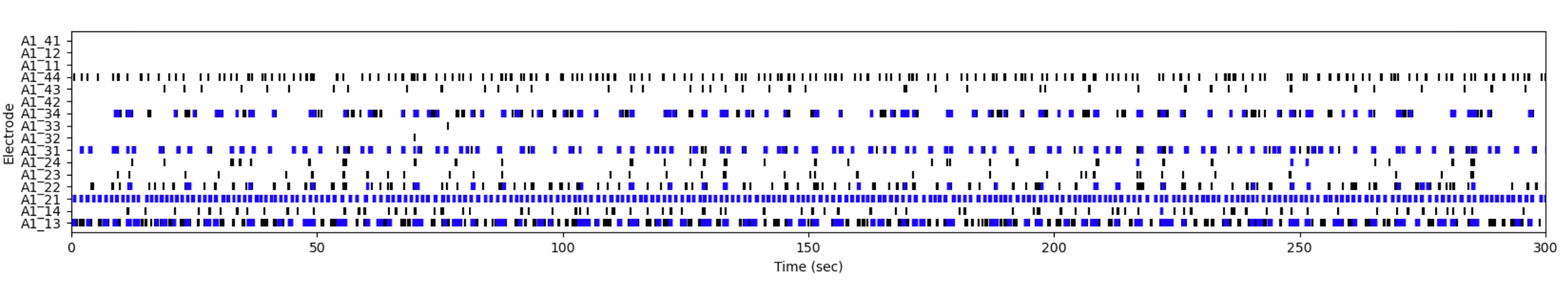
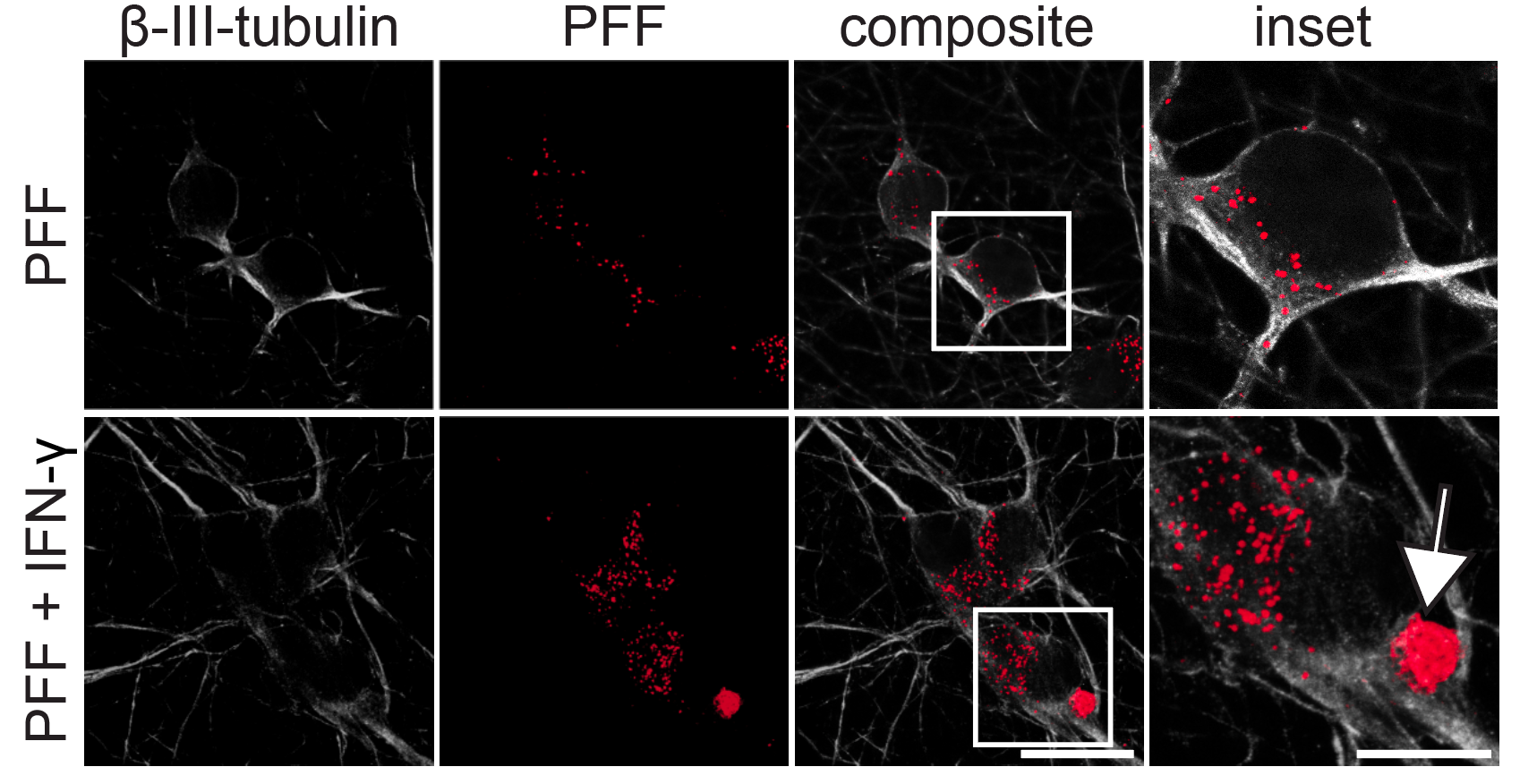
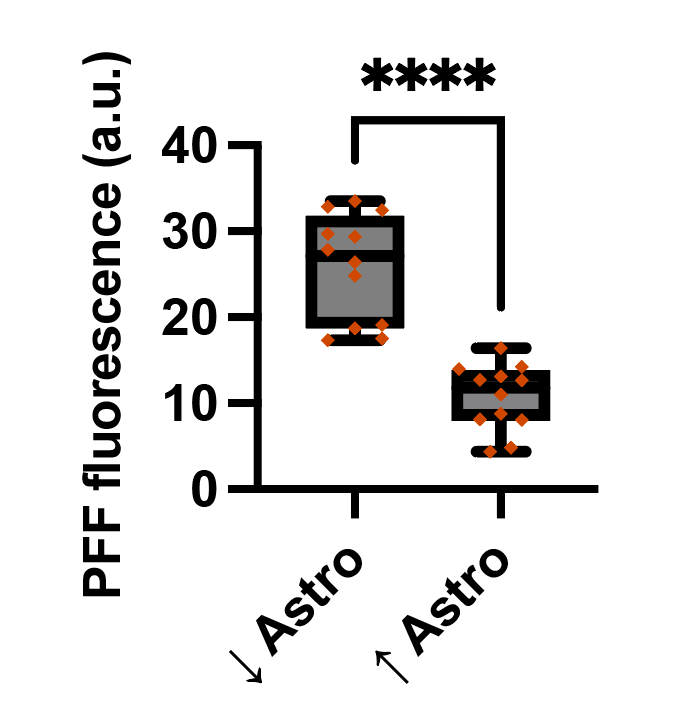
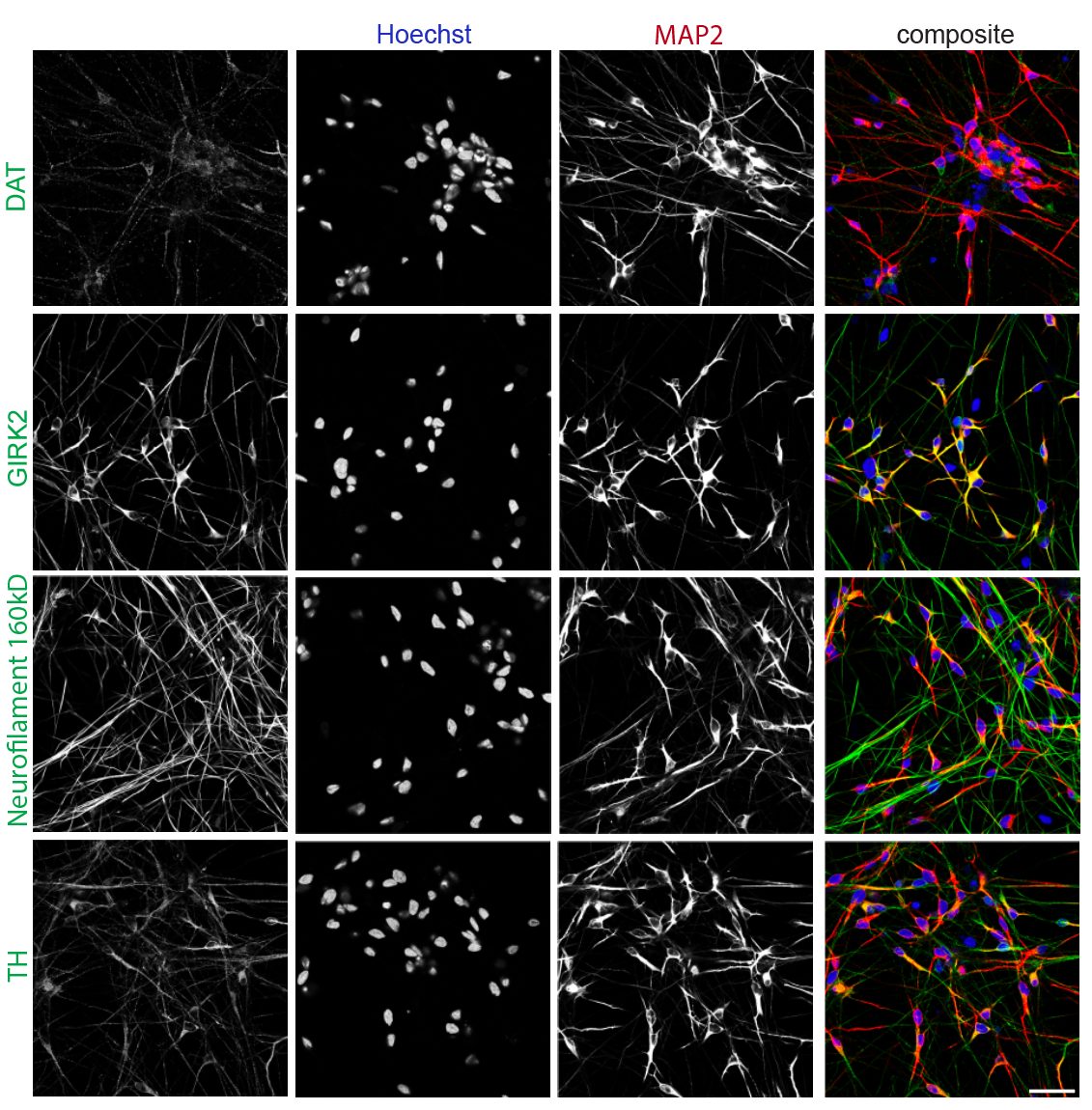
Following differentiation, I characterize neuronal identity via antibody profiling (ICC), qPCR, and flow cytometry. Electrical activity is confirmed using microelectrode arrays (MEAs) for spontaneous firing patterns and waveform analysis. I have also performed HPLC-based quantification of dopamine and metabolites from culture media to link biochemical outputs to functional phenotypes.
RT-qPCR quantification confirms cell-type identity and treatment response at the transcriptional level. Expression is normalized to housekeeping genes (GAPDH, ACTB) and presented as fold change relative to undifferentiated or untreated controls. Error bars represent SEM from n=3 biological replicates.
I confirm neuronal identity and functional maturity using microelectrode array (MEA) recordings and calcium imaging. MEA captures spontaneous electrical activity, burst frequency, and network synchrony across neuronal subtypes. Calcium imaging with Fluo-4 or GCaMP reporters reveals spontaneous and evoked calcium transients in real time.