Movement Disorders — An Integrated Research Program
My work centers on the molecular and cellular biology of Parkinson's disease — α-synuclein pathology, lysosomal dysfunction, dopaminergic neuron vulnerability, and neuroinflammation — built on iPSC-derived human neuron platforms, CRISPR-edited disease lines, and quantitative multi-modal readouts. The same platform translates directly to dystonia and other basal-ganglia / motor-circuit disorders: iPSC differentiation into dopaminergic and cortical/striatal-projecting neurons, microelectrode-array (MEA) electrophysiology, calcium imaging, high-content confocal and electron microscopy, CRISPR knockout modeling, and mass-spectrometry proteomics. This page consolidates the core figures, assays, and findings from across the site that are directly relevant to evaluating my capabilities in Parkinson's disease and dystonia.
Research Pillars
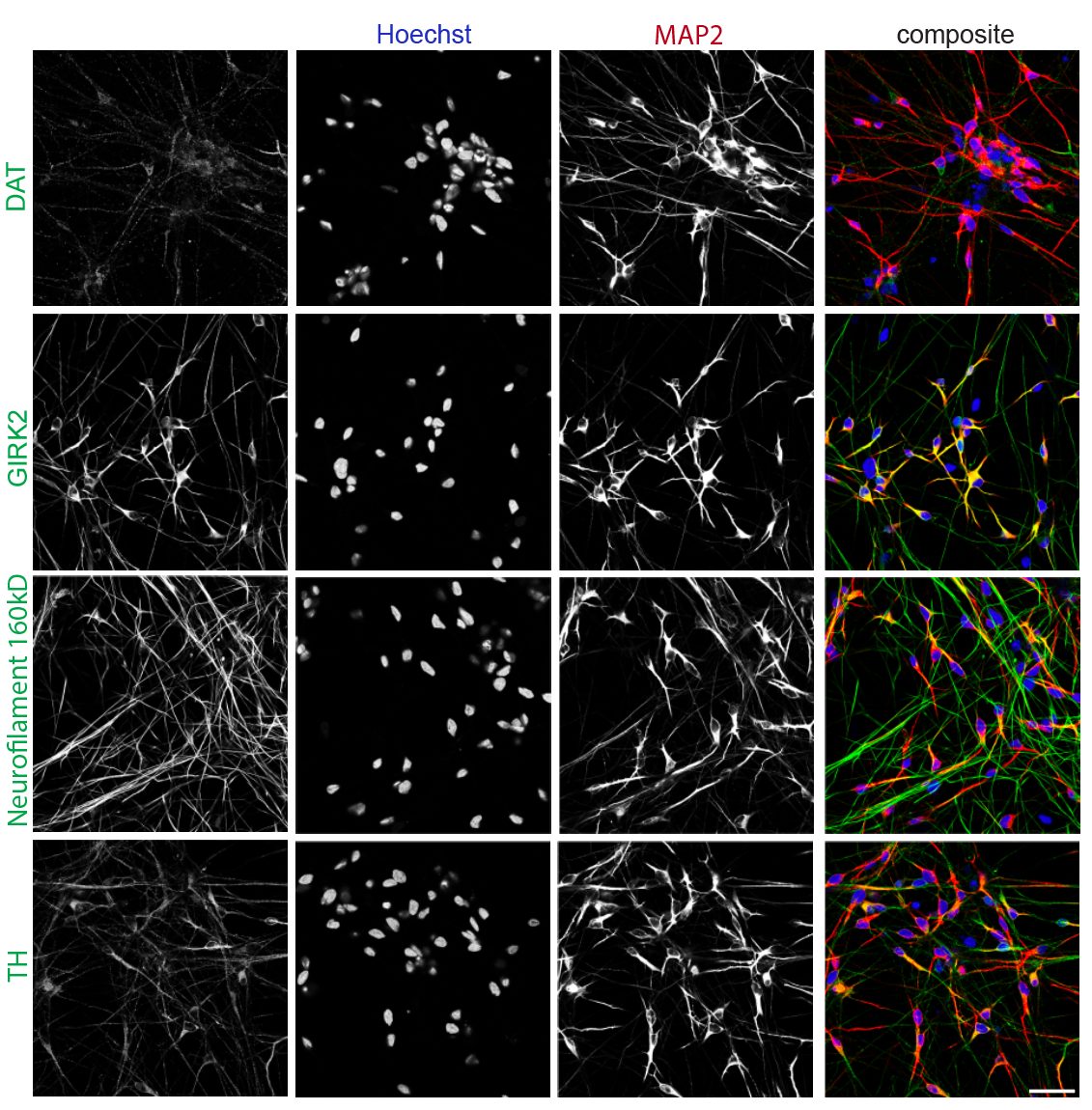
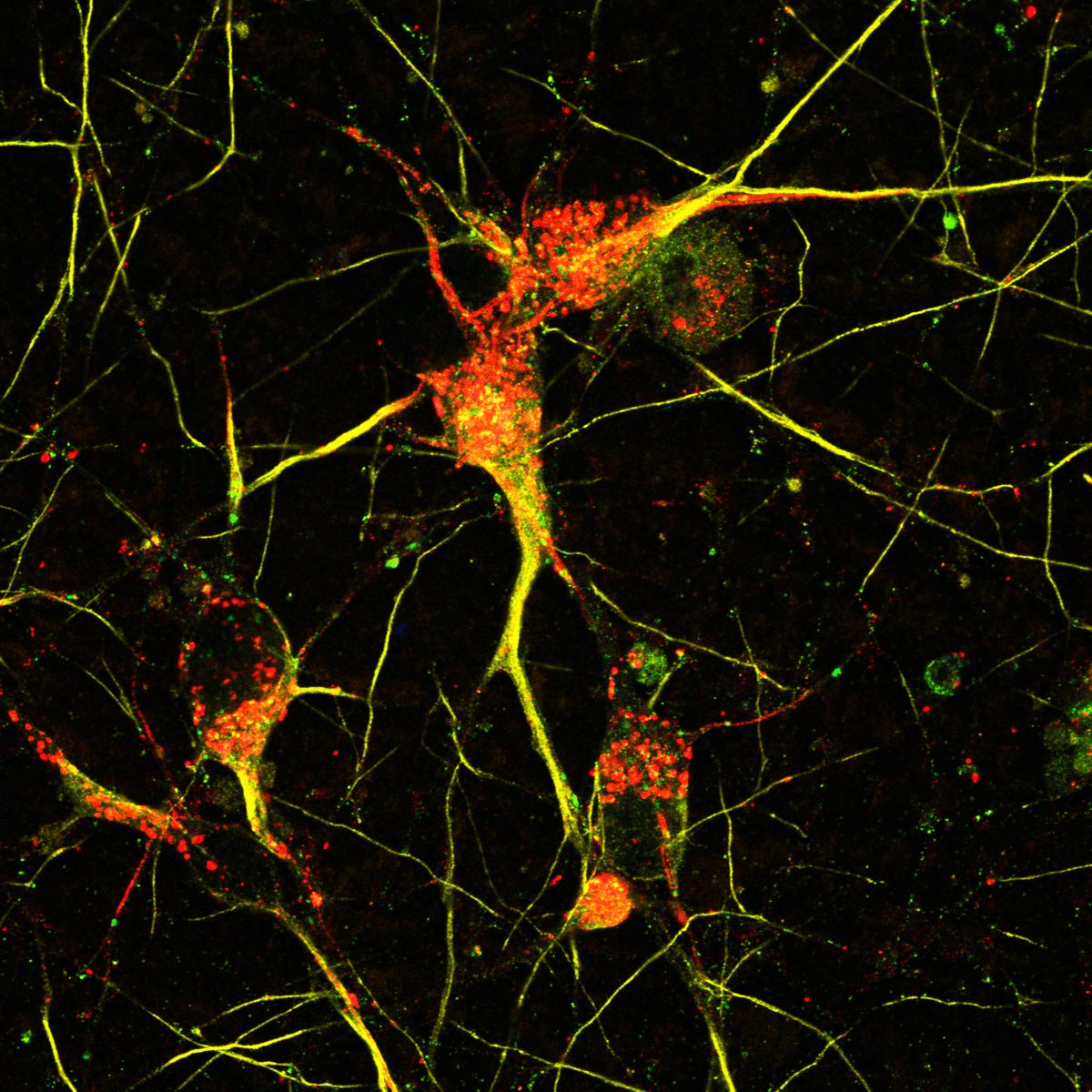
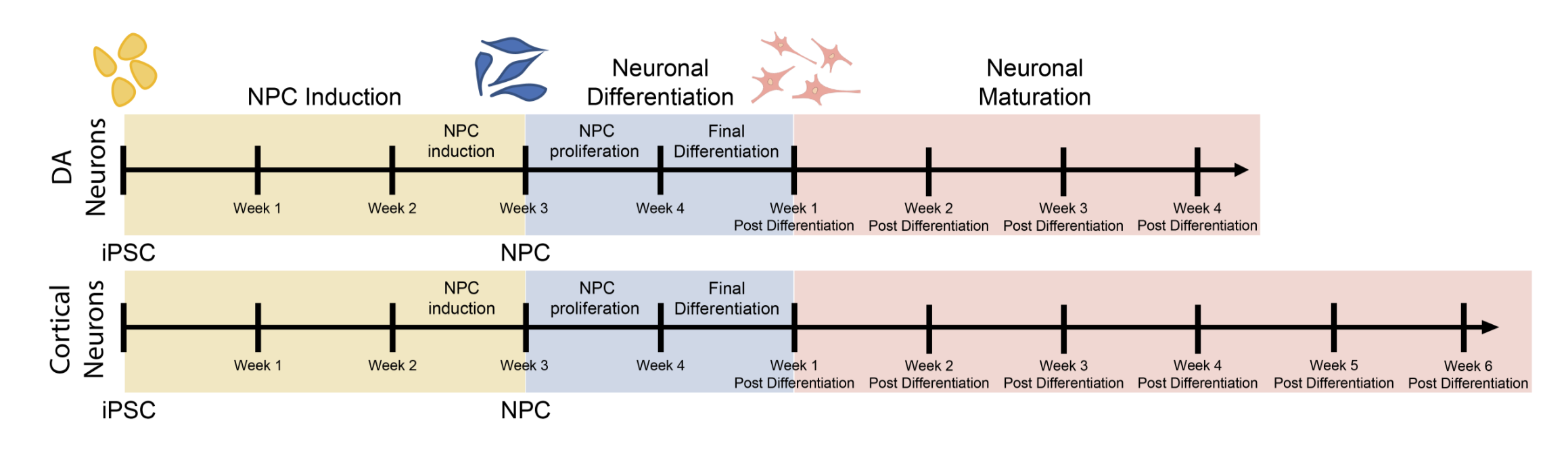
- iPSC-derived midbrain dopaminergic & cortical neurons
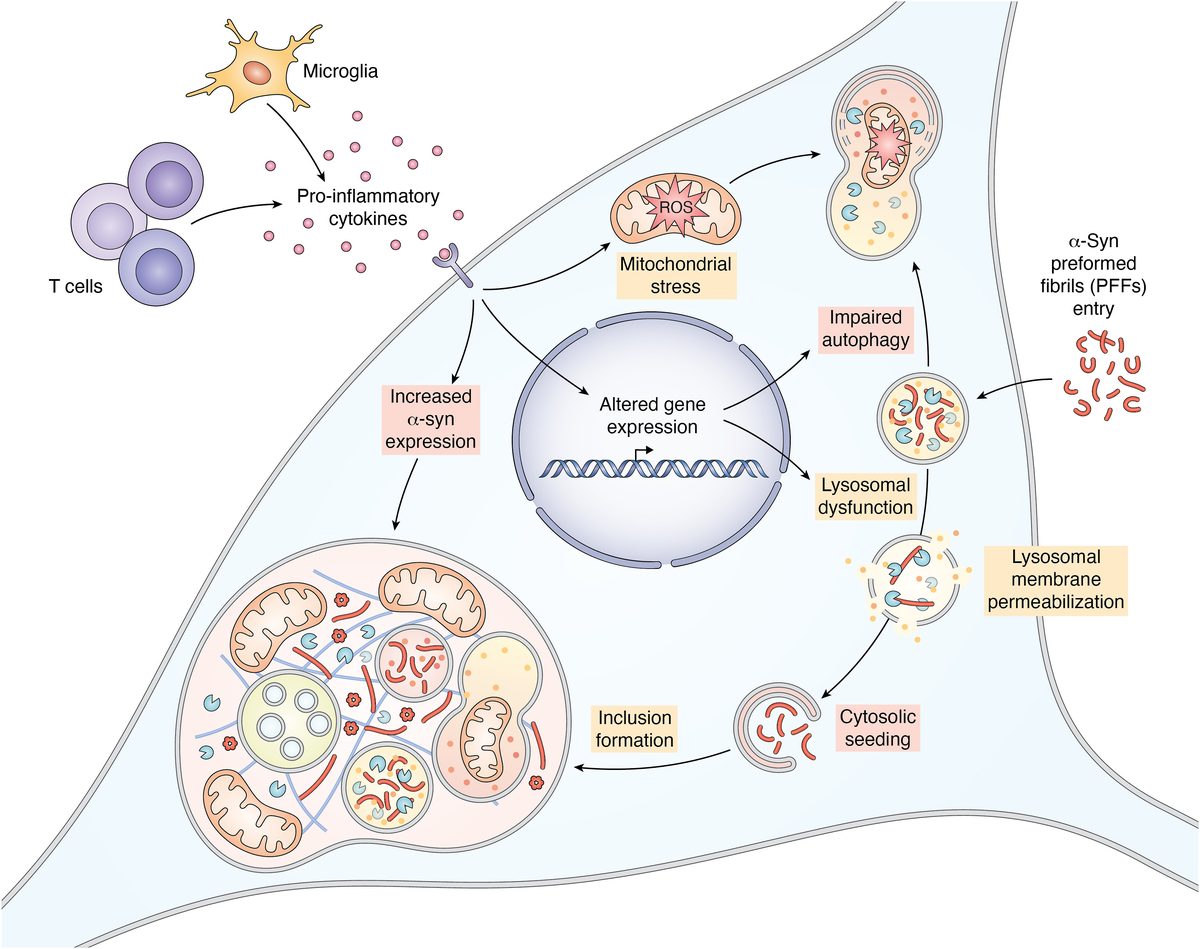
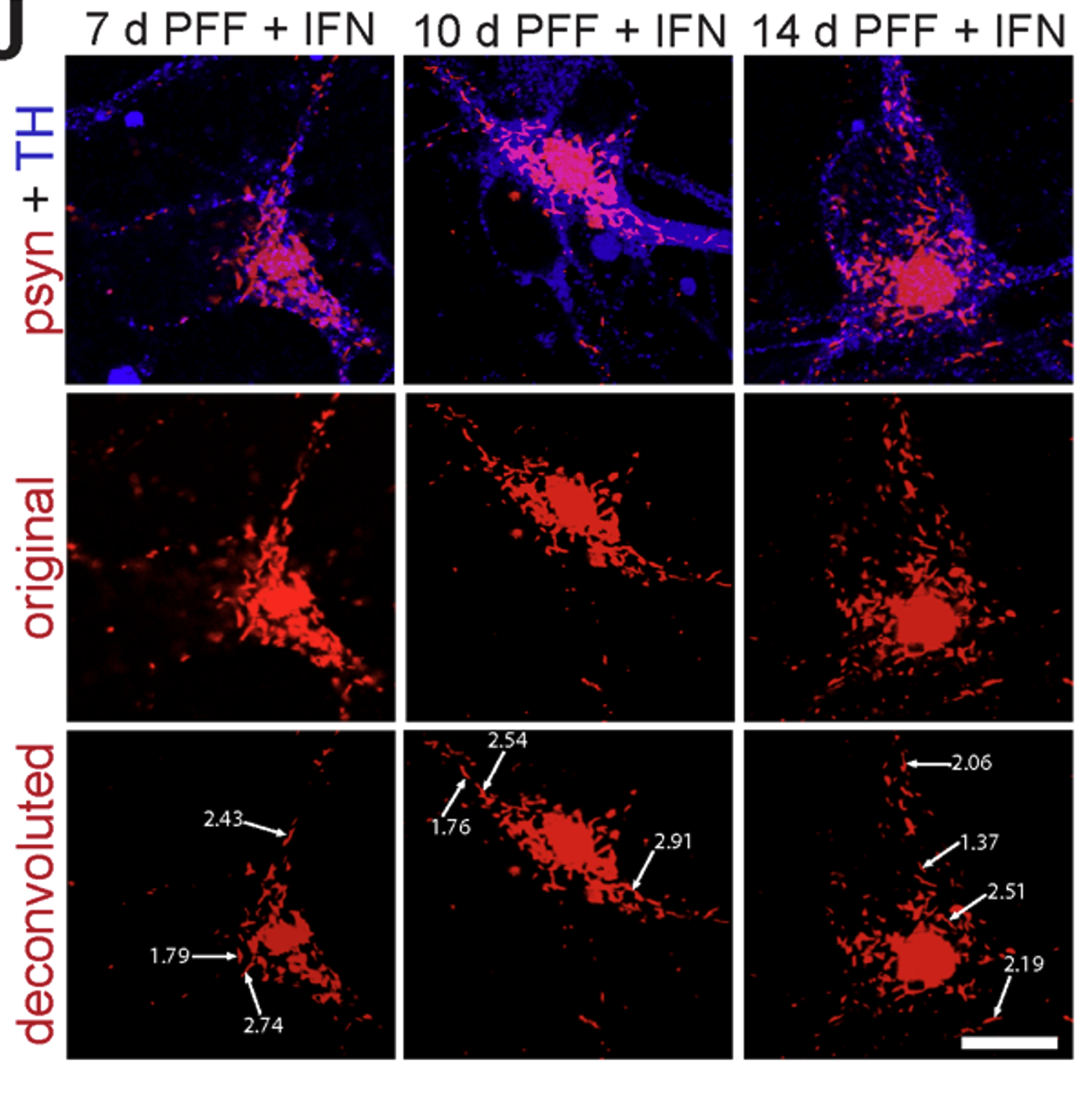
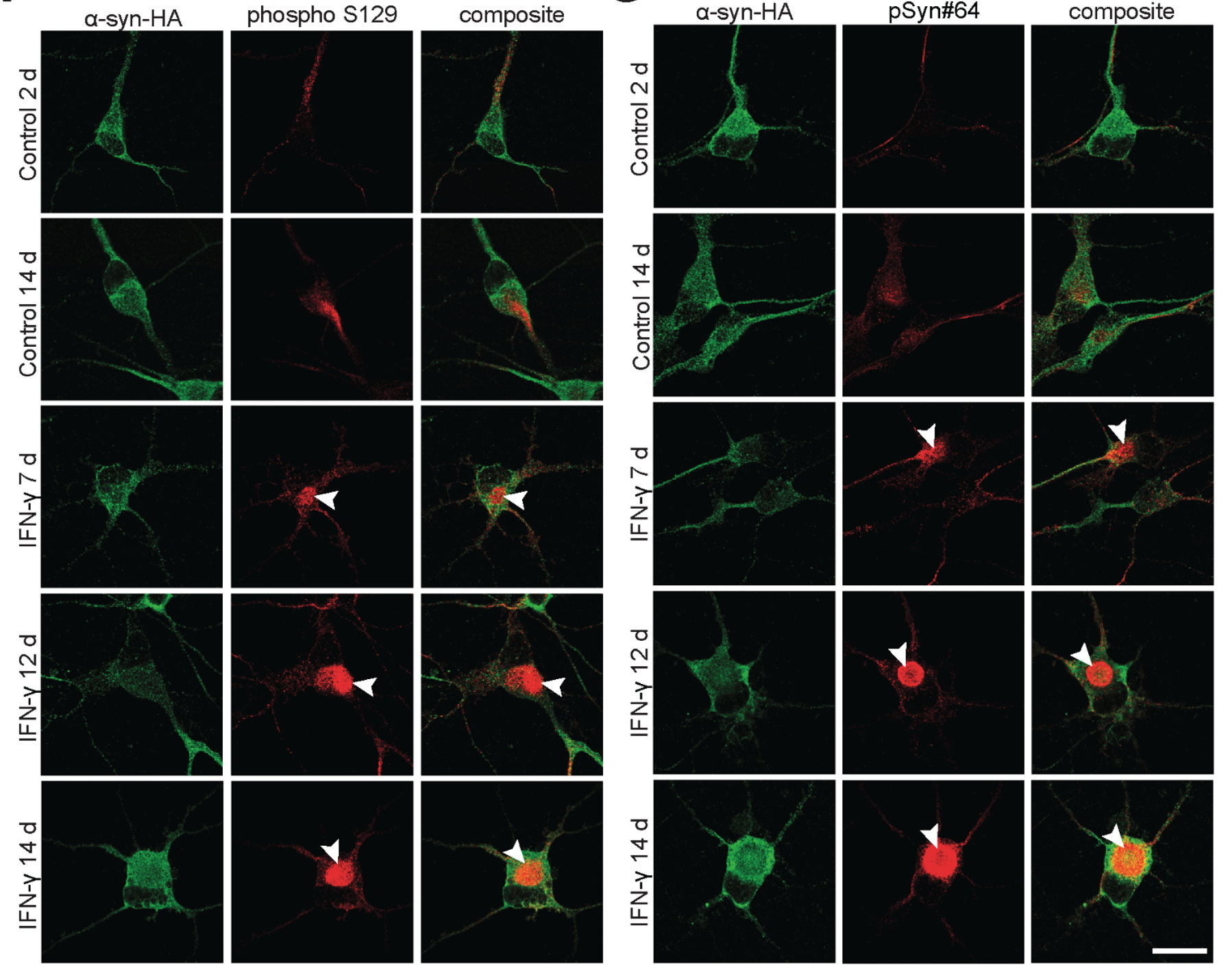
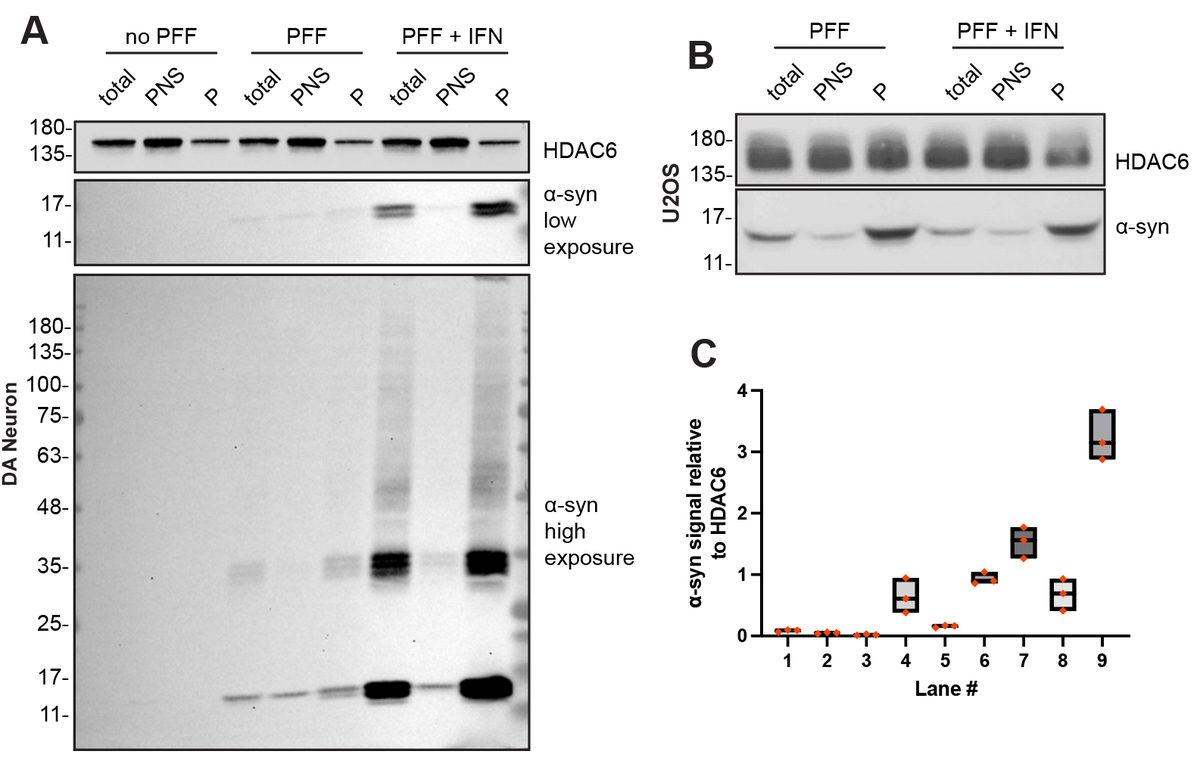
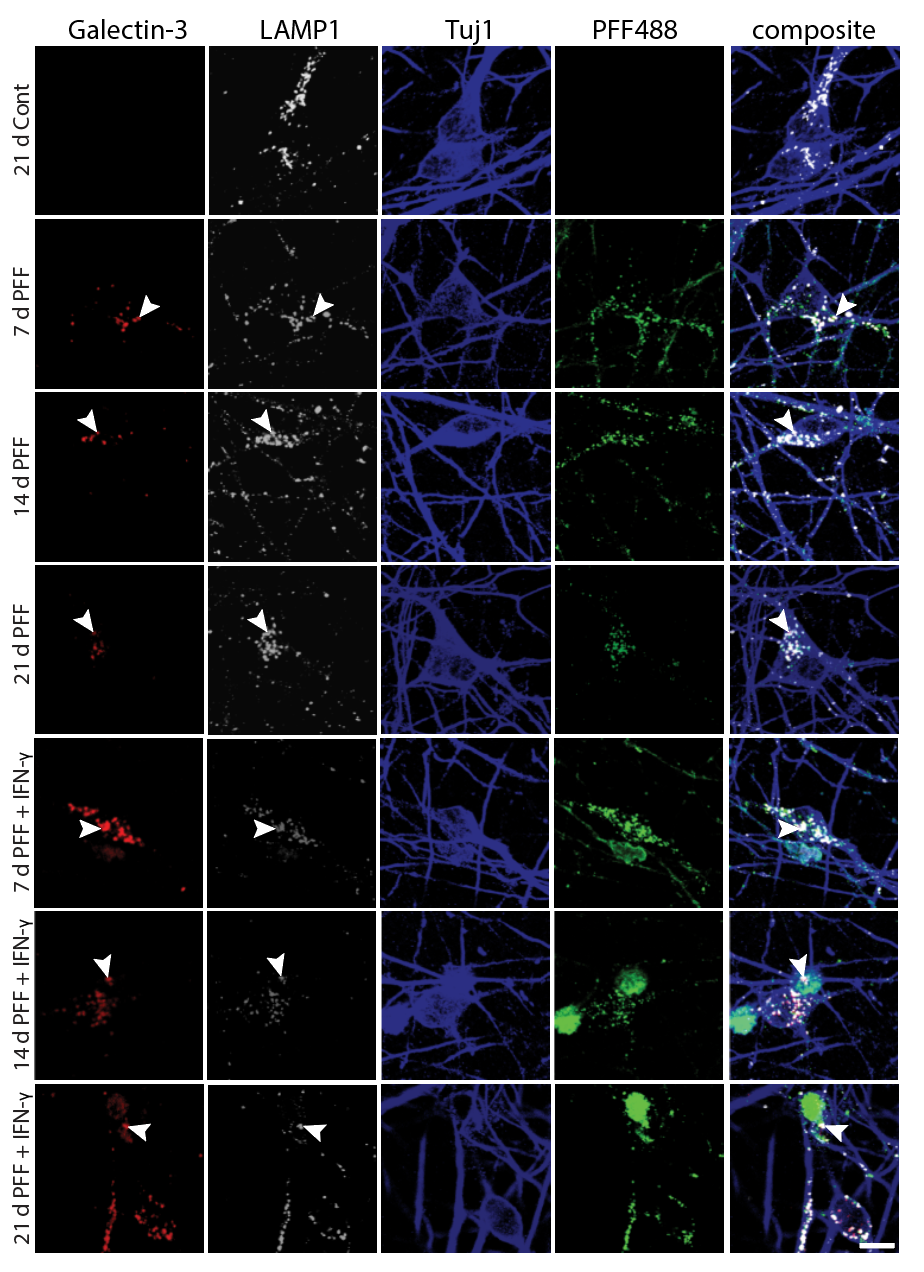
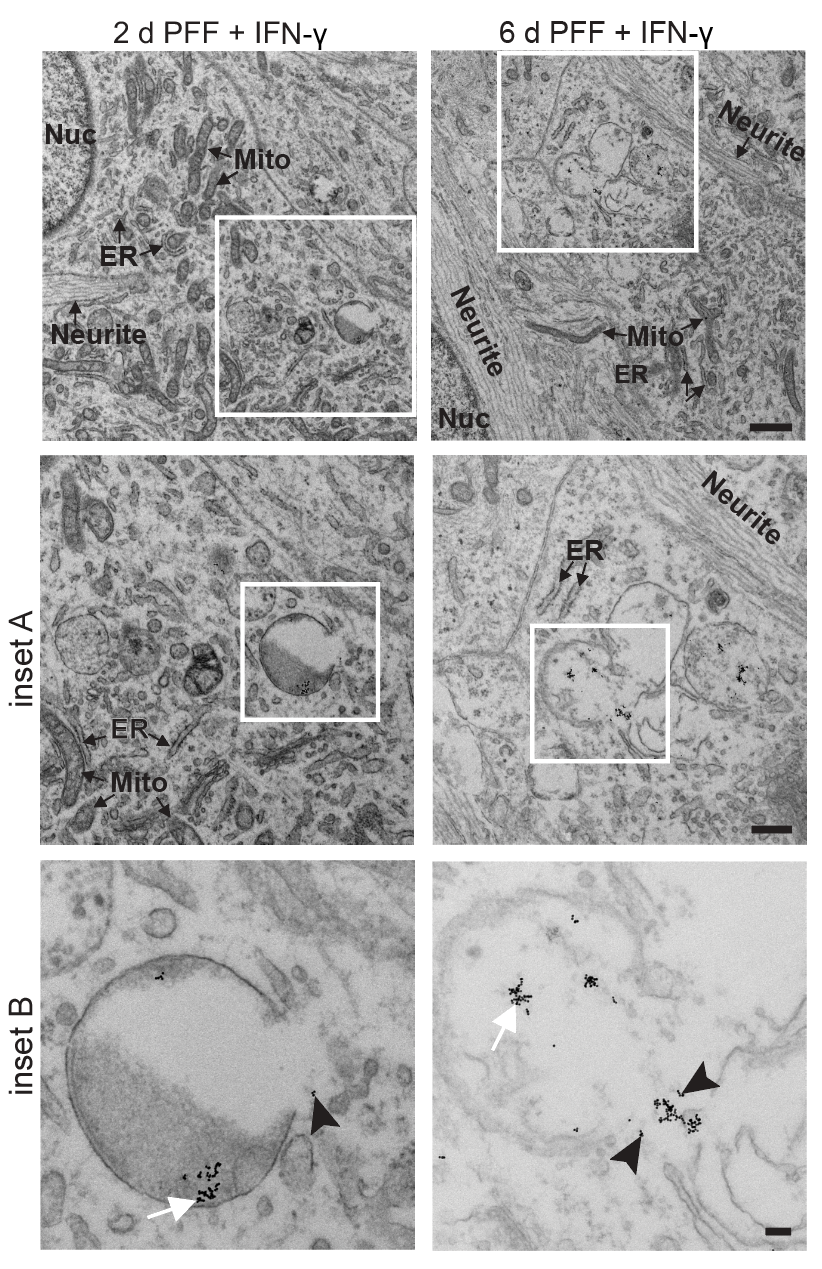
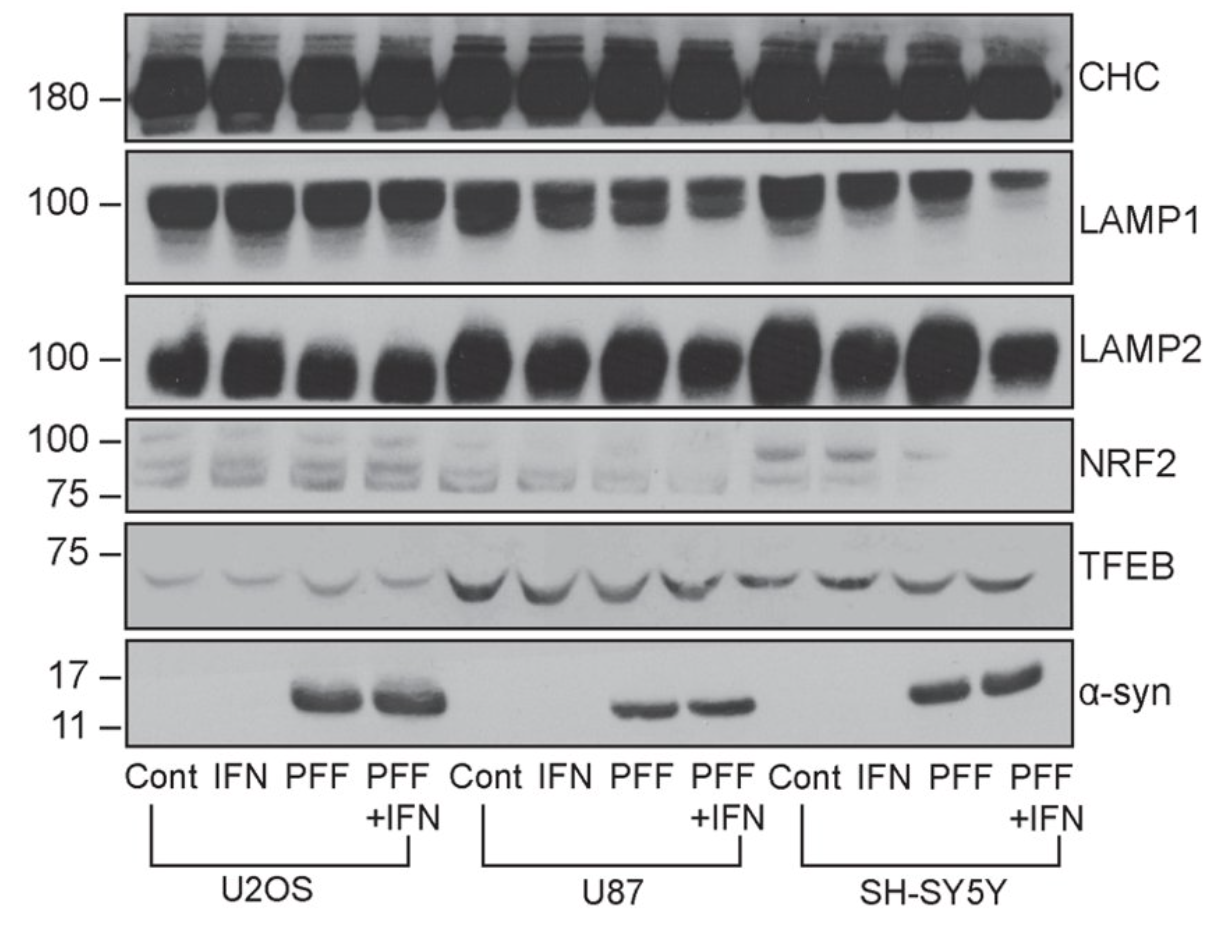
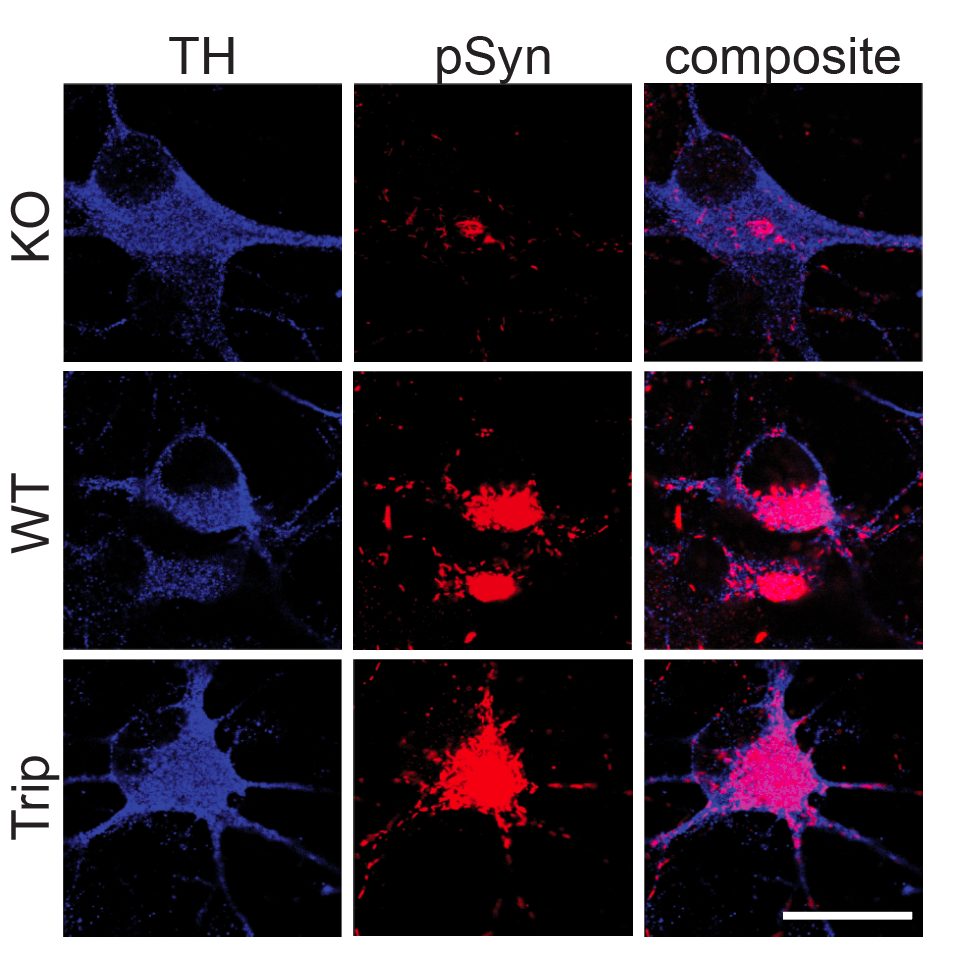
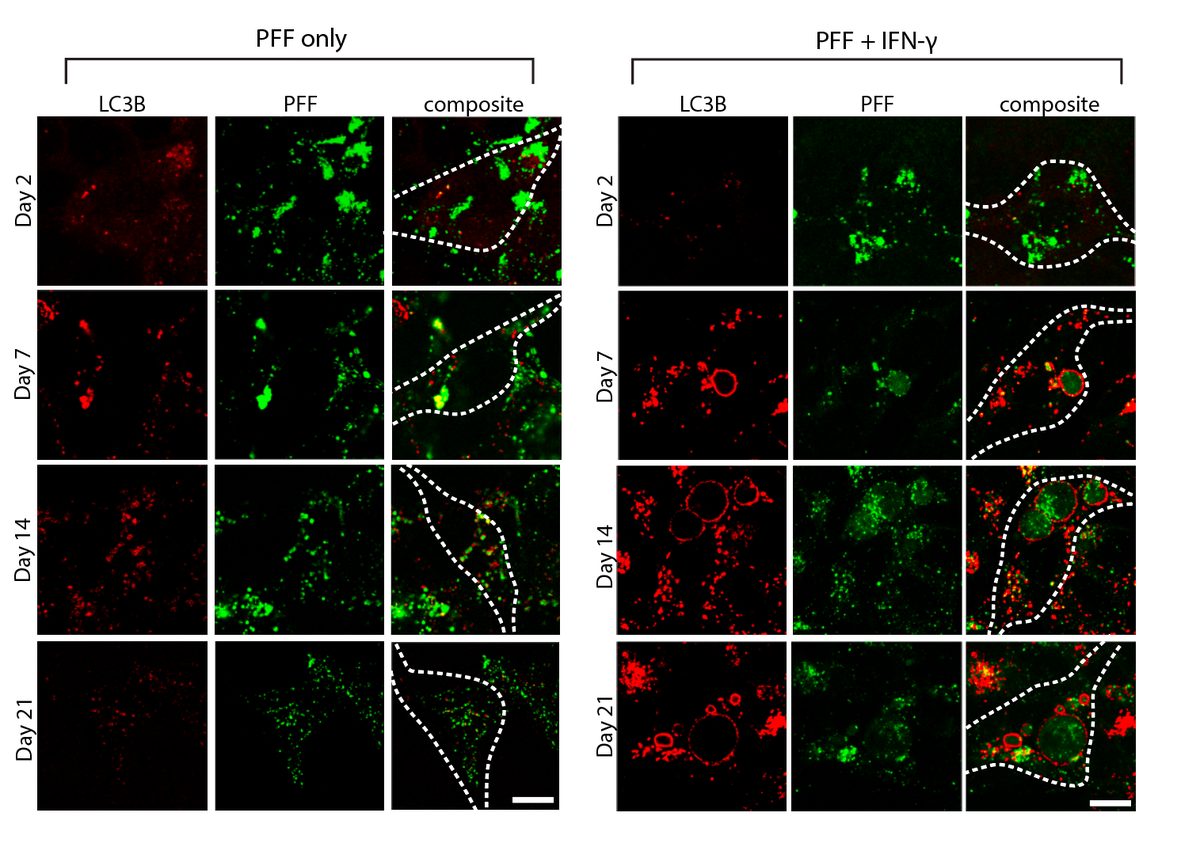
- α-Synuclein PFF seeding & dual-hit Lewy-body modeling
- CRISPR-edited disease lines (LRRK2, GBA, Parkin, SNCA)
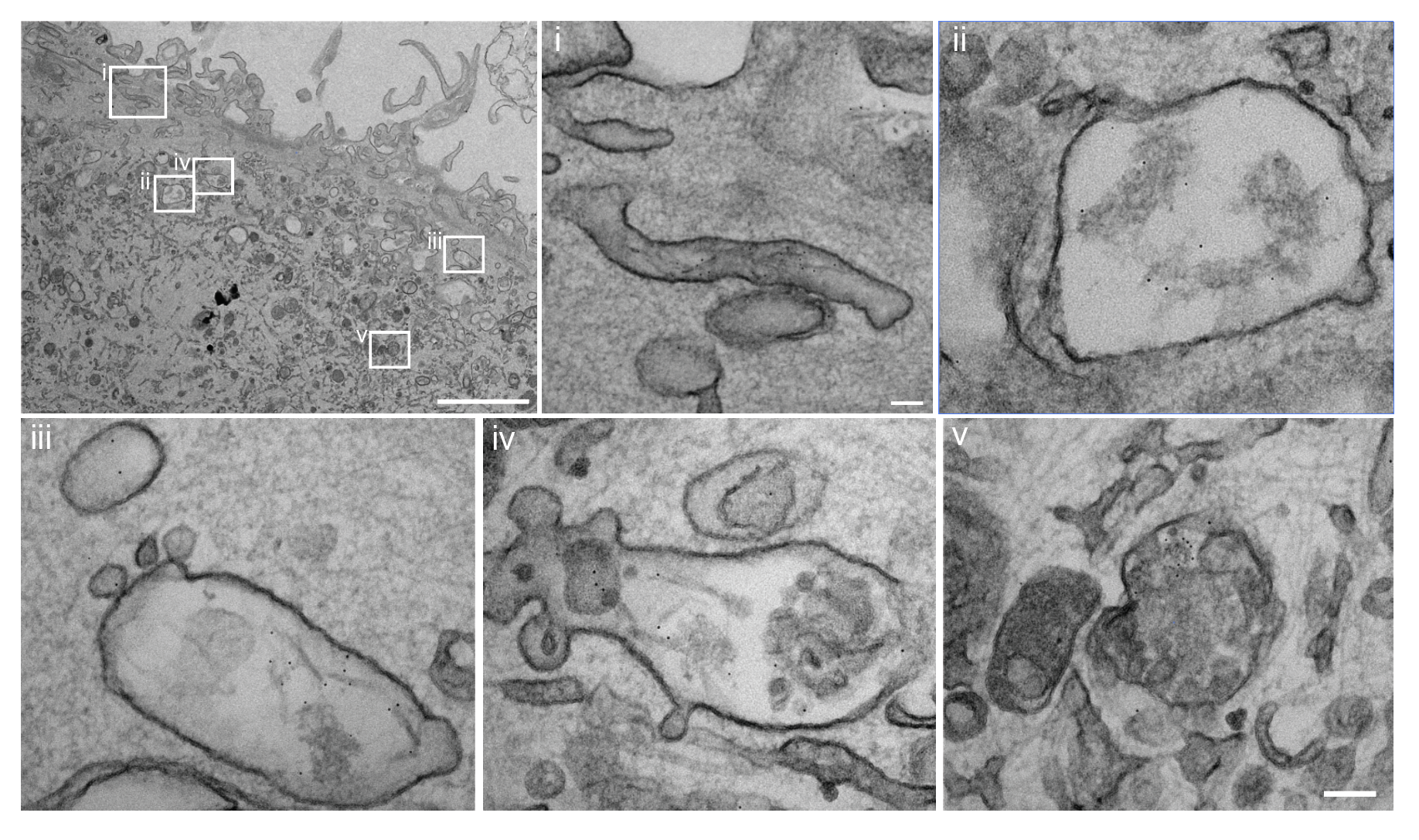
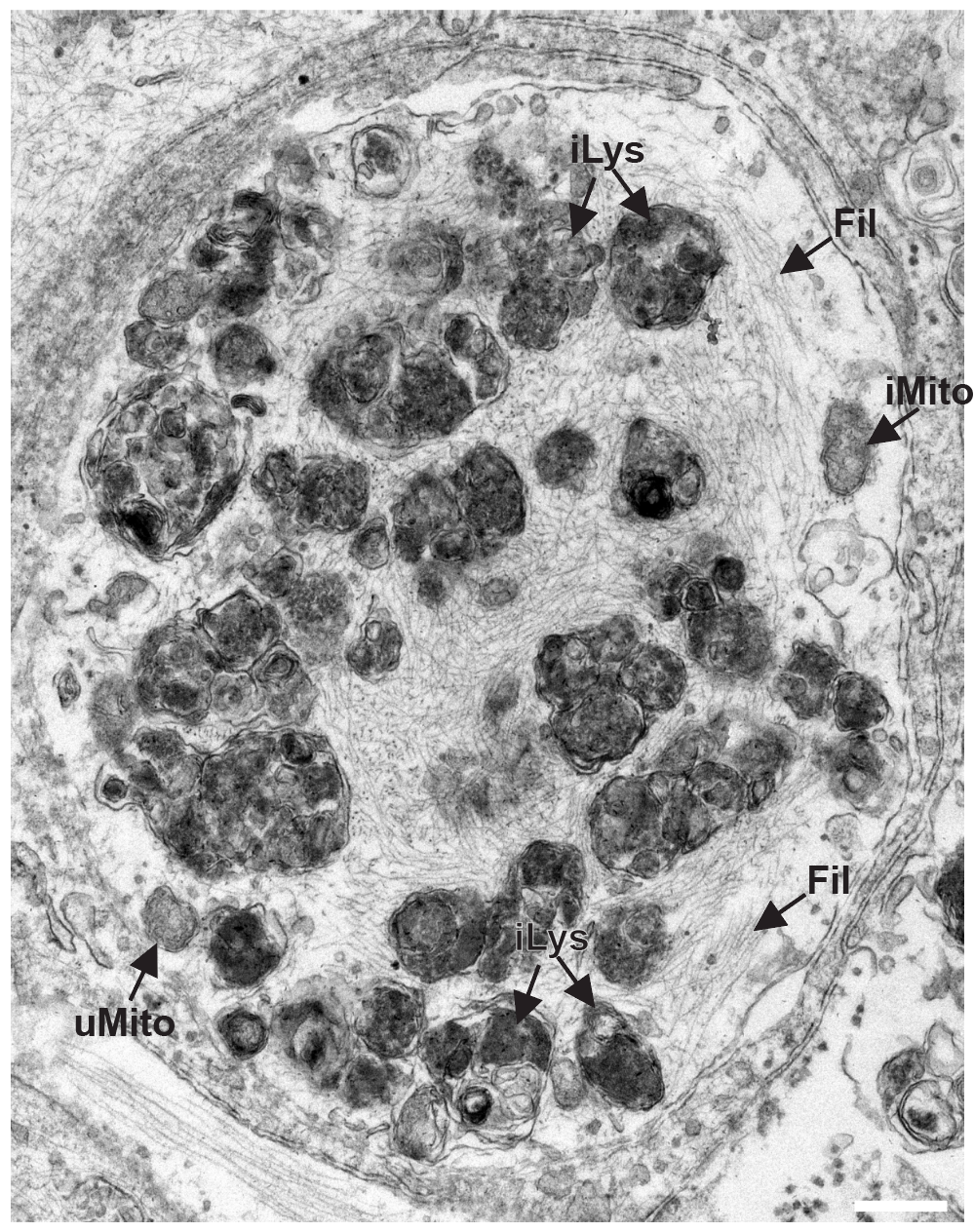
- Lysosomal dysfunction & membrane permeabilization
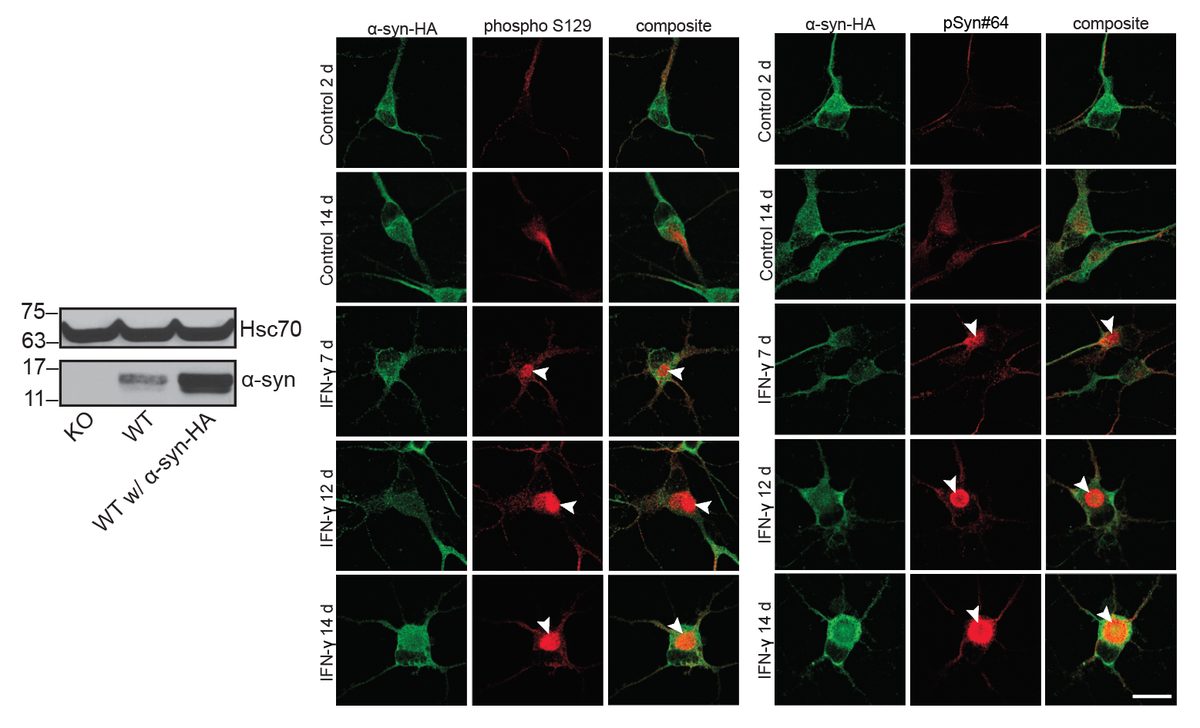
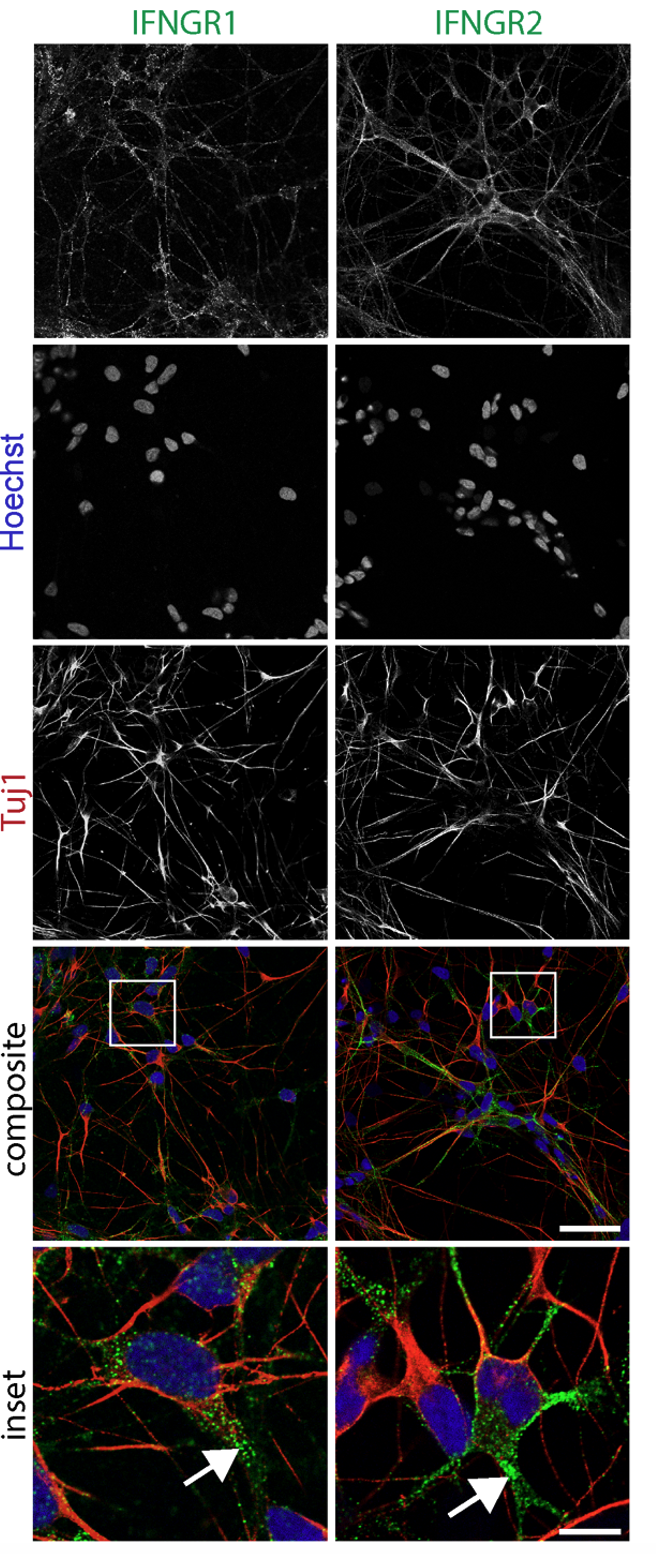
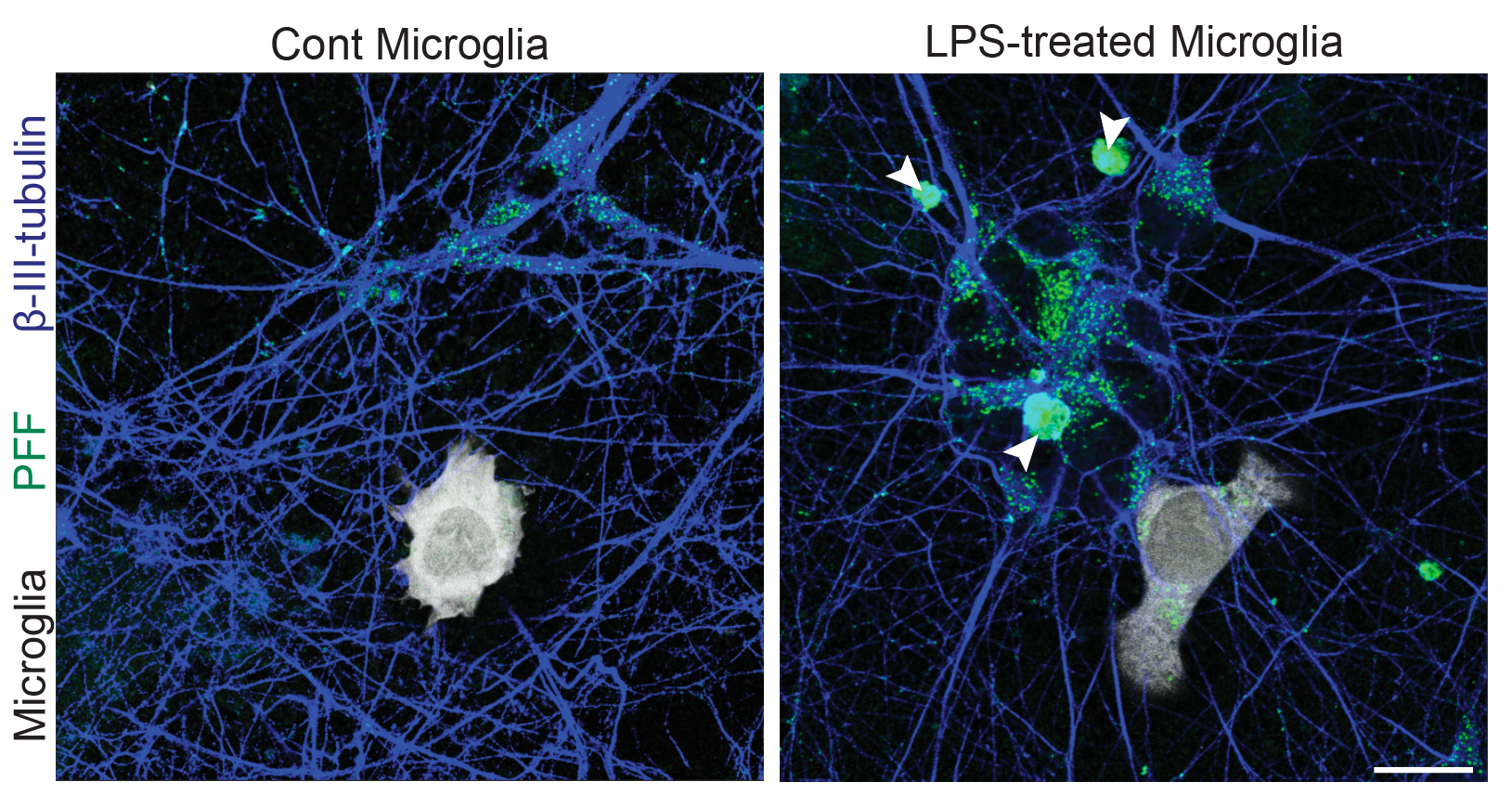
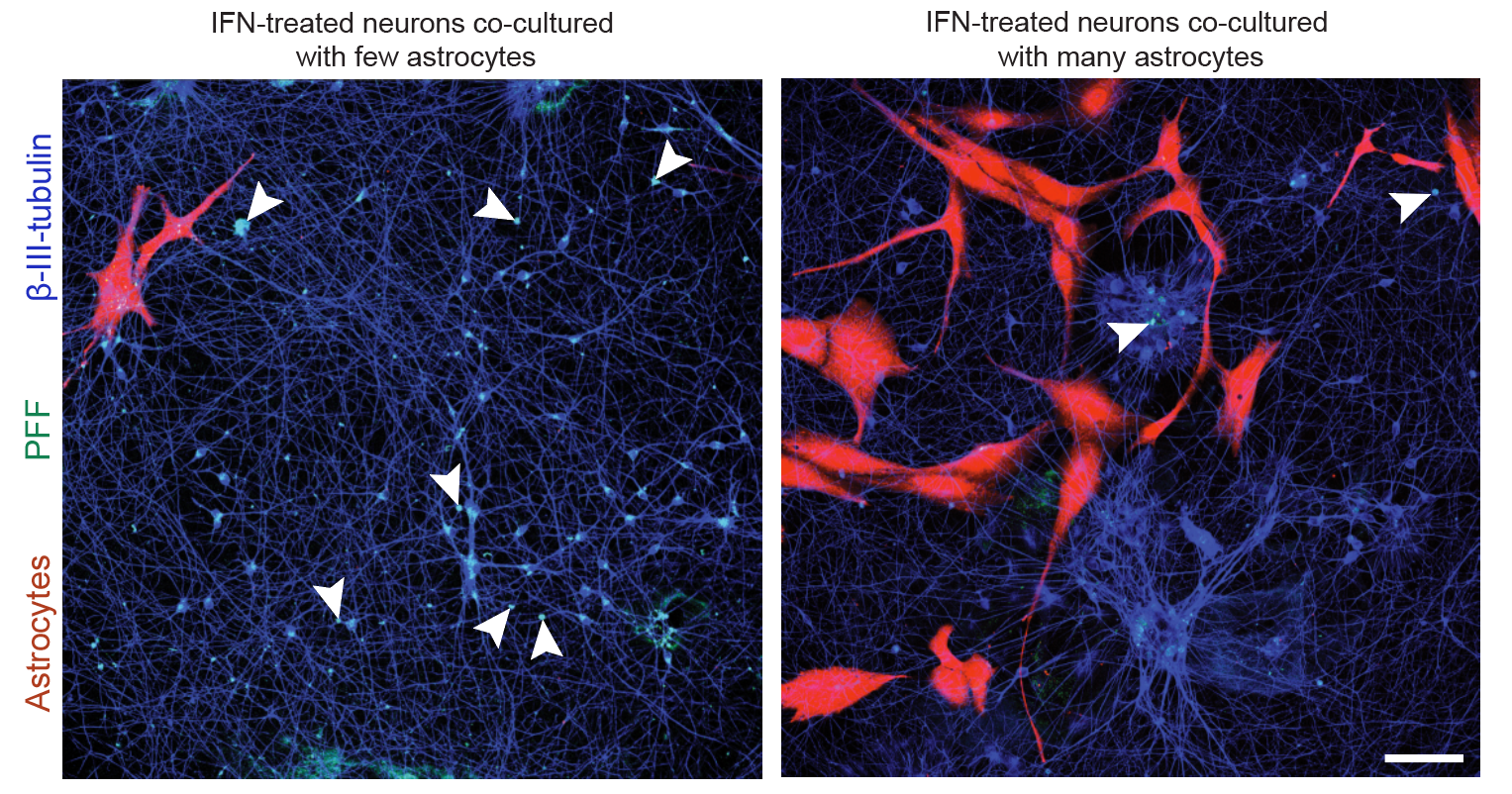
- Neuroinflammation (IFN-γ, TNF-α, IL-1β) and microglial co-culture
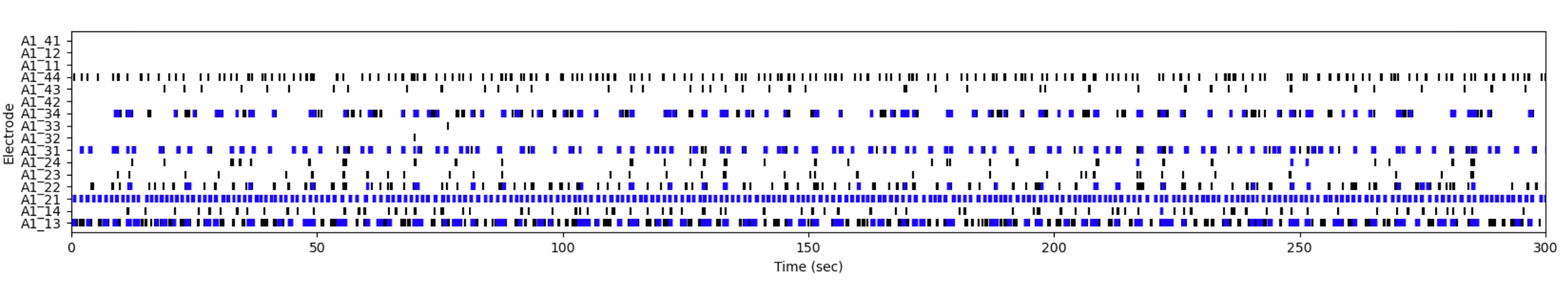
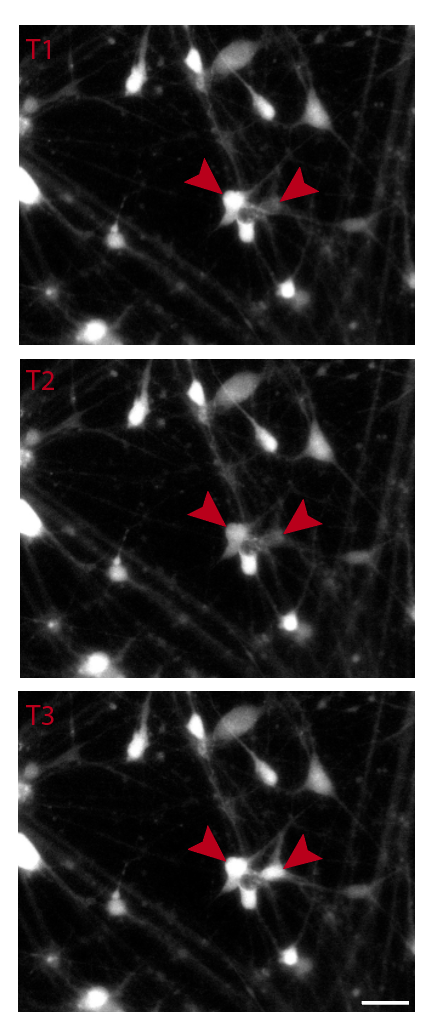
- MEA electrophysiology & calcium imaging — circuit-level readouts for movement disorders
- Parkin-agonist & lysosomal-rescue small-molecule discovery
- Biomarker development: pS129, LAMP1/2, TFEB, NfL, HDAC6